Most currently-available vaccines were empirically designed, and contain attenuated or killed pathogens. The need to cultivate infectious agents and isolate the inactivated whole pathogen or some of its purified components, as well as concerns of safety, rendered this approach elusive for many pathogens. As part of the IIBR ongoing effort to develop novel countermeasures against bio-threat agents, we are establishing state-of-the-art novel strategies for the rational identification of immune-related components, towards the design of novel vaccines. The availability of complete genomic sequence of various pathogens revolutionized the field of vaccinology.
Concomitant with the development of in silico genome mining procedures and high-throughput global technologies, the paradigm of vaccine development shifted in recent years from conventional culture-based methods to high throughput approaches, based on in silico analyses of the entire repertoire of the pathogen’s genome-encoded genes, followed by extensive experimental assessment, aiming at faster and more comprehensive identification of novel targets for attenuation and/or design of subunit vaccines.
At the same time, identification of the vital role of cellular immunity in defense against intra-cellular pathogens resulted in incorporation of the cellular immunity aspects into reverse vaccinology, referred to as ‘‘reverse immunology’’. A line of advanced, highly-reliable computational tools were developed in recent years for mapping the potential repertoire of T-cell epitopes for a given pathogen. The IIBR is developing novel, unbiased whole-genome-based strategies for rational down selection of potential candidates, as well as platforms for high throughput experimental evaluation of their protective immunity, and the institute is pioneering global immunoinformatic screens of proteomes of complex pathogens.
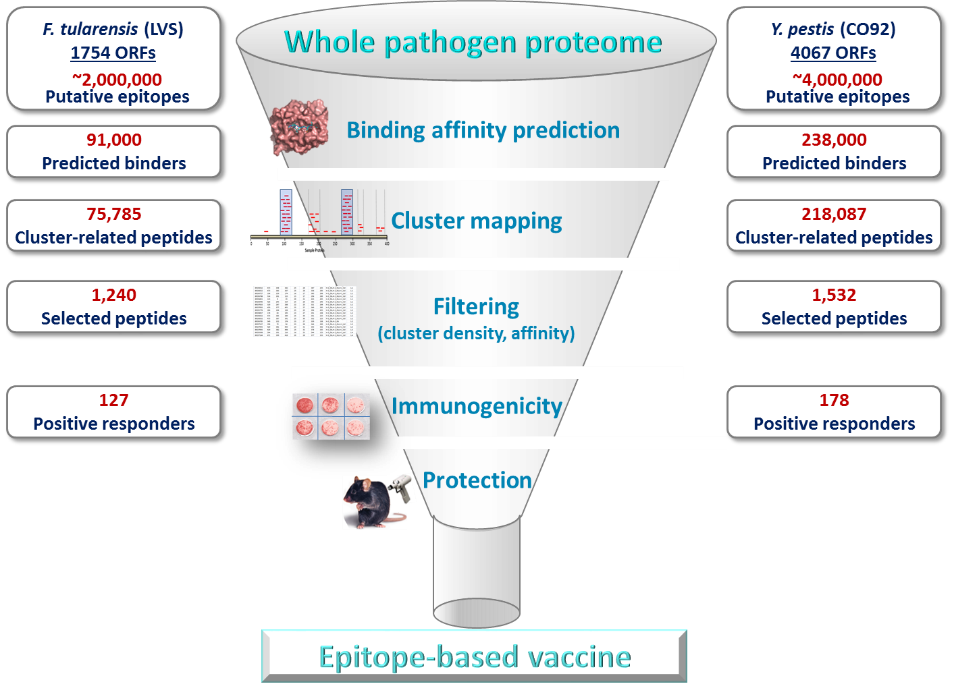
Illustration of the whole-genome based combined in silico and experimental multi-step strategy for identification of novel protective epitopes